Link: Multiple regression R Linear regression in R
Check the full code in github repo.
Do multiple regression in R
Plot the basic correlation
library(dplyr)
mouse.data <- data.frame(
size = c(1.4, 2.6, 1.0, 3.7, 5.5, 3.2, 3.0, 4.9, 6.3),
weight = c(0.9, 1.8, 2.4, 3.5, 3.9, 4.4, 5.1, 5.6, 6.3),
tail = c(0.7, 1.3, 0.7, 2.0, 3.6, 3.0, 2.9, 3.9, 4.0))
plot(mouse.data)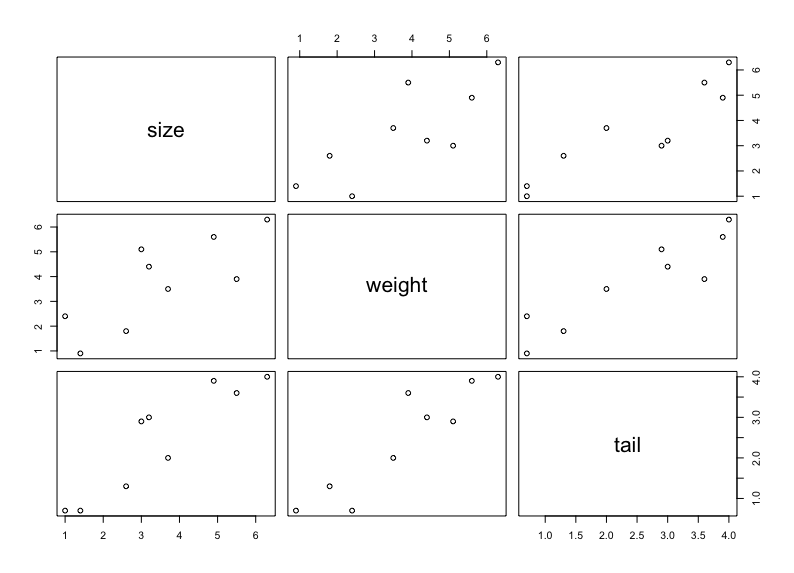
Interpret the graph
Identify the correlation by eye-balling:
- Both weight and tail are correlated with size.
- weight and tail are also correlated - so we might not need both
Fit a plane to the data
Use ”+” sign in lm() to indicate multiple parameters.
multiple.regression <- lm(size ~ weight + tail, data=mouse.data)Use summary() to interpret the results
Note: since we are using multiple regression, the adjusted R-squared would be more meaningful.
Call:
lm(formula = size ~ weight + tail, data = mouse.data)
Residuals:
Min 1Q Median 3Q Max
-0.99928 -0.38648 -0.06967 0.34454 1.07932
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.7070 0.6510 1.086 0.3192
weight -0.3293 0.3933 -0.837 0.4345
tail 1.6470 0.5363 3.071 0.0219 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.8017 on 6 degrees of freedom
Multiple R-squared: 0.8496, Adjusted R-squared: 0.7995
F-statistic: 16.95 on 2 and 6 DF, p-value: 0.003399How to compare between multiple vs linear regression
We can do so by comparing the coefficient table with simple linear regression:
The lines of x variables represents the results of multipe regression while comparing with simple linear regression without it.
e.g. The weight line compares the multipel regression y = n + a*weight + b*tail vs simple regression y = n + b*tail. In this case, p-value > 0.05, so weight + tail is not significantly better(important) than using tail alone to predict size, meaning we can get rid of weight in the model.